Convocatoria 2022 - «Proyectos de Generación de Conocimiento»
The scientific community reacted to the COVID-19 pandemic in an outstandingly rapid and collaborative way. We now have more information on a virus than ever before thanks to the publication of thousands of articles and related data after three years of research on this subject. This information has been used to develop effective vaccines and antivirals drugs against COVID-19. To combat emerging SARS-CoV-2 variants that may be resistant to current treatments or novel viruses that may trigger future pandemics, research in this field must be continued. For this reason, we propose this three-year project submitted to the research-oriented modality and related to the most important thematic priority in health that we have had in recent years. This is a multidisciplinary project that brings together a group of computer scientists and chemists with the aim of generating new computational tools that will help the scientific community to search for antiviral drugs against current and future pandemics.
Desarrollo de nuevas herramientas computacionales para el descubrimiento de fármacos contra las pandemias actuales y futuras (NEXT-PANDEMICS) (PID2022-138327OB-I00)
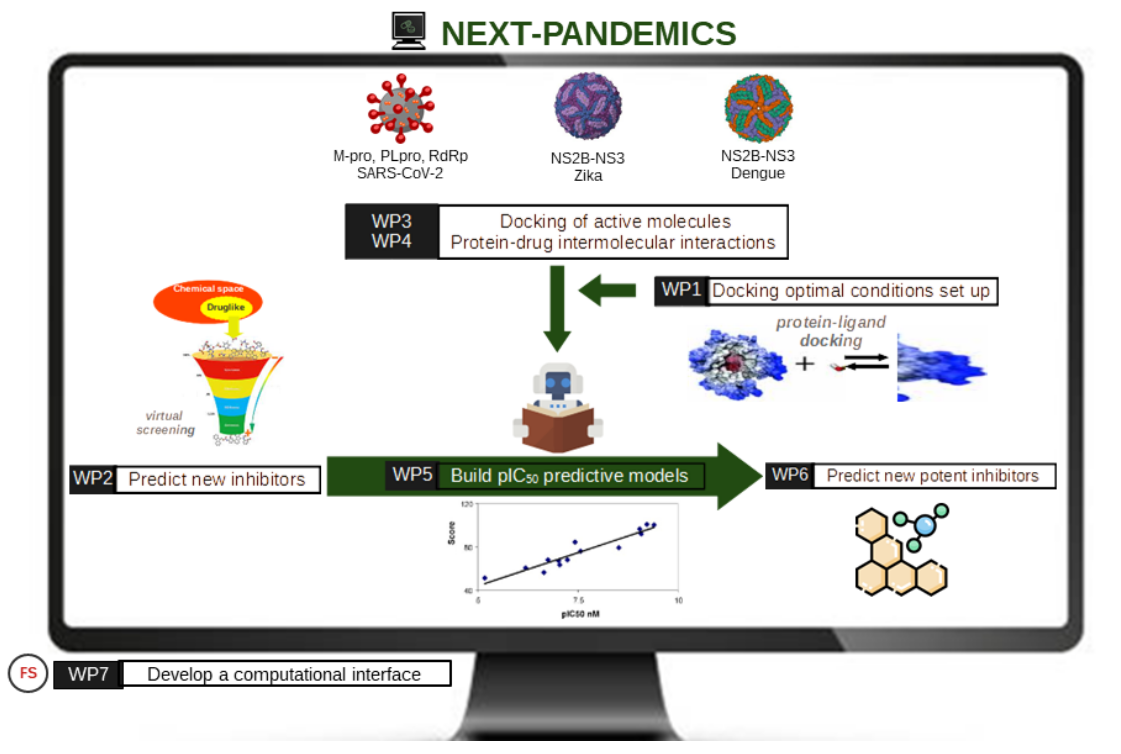 Next-pandemics
Next-pandemics
Team:
- Santiago Garcia-Vallvé, PhD. Cheminformatics and Nutrition Research Group. Biochemistry and Biotechnology Department. Rovira i Virgili University (URV)
- Francesc Serratosa Casanelles, PhD. Department of Computer Engineering and Mathematics (DEIM). Rovira i Virgili University (URV)
- Gerard Pujadas Anguiano, PhD. Cheminformatics and Nutrition Research Group. Biochemistry and Biotechnology Department. Rovira i Virgili University (URV)
- Aleix Gimeno Vives, PhD. Cheminformatics and Nutrition Research Group. Biochemistry and Biotechnology Department. Rovira i Virgili University (URV)
- Susana Álvarez Fernández, PhD. Department of Computer Engineering and Mathematics (DEIM). Rovira i Virgili University (URV)
- Maria Carmen Julià Ferré, PhD. Department of Computer Engineering and Mathematics (DEIM). Rovira i Virgili University (URV)
- Ariadna Llop Peiró, PhD student. Cheminformatics and Nutrition Research Group. Biochemistry and Biotechnology Department. Rovira i Virgili University (URV)
- Sarah Fadlallah, PhD student. Department of Computer Engineering and Mathematics (DEIM). Rovira i Virgili University (URV)
- Natàlia Segura Alabart, PhD student. Department of Computer Engineering and Mathematics (DEIM). Rovira i Virgili University (URV)
Results:
-
Llop-Peiró A, Sun L. Prediction of Aryl Hydrocarbon Receptor (AHR) inhibitors: Dry lab meets wet lab. 15th Annual Harvard Surgery Research Day. Boston, USA. 13th March 2026.
-
Llop-Peiró A, Silva GM, Sun L. Computational Insights into Dual-specificity Tyrosine Phosphorylation-regulated Kinase 1A (DYRK1A) Selectivity. OpenEye CUP. Sanfa Fe, New Mexico, USA. 10-12 March 2026.
-
Trujillo-De León S, Llop-Peiró A, Pujadas G, Garcia-Vallvé S, Gimeno A. pICkIT: Automated Analysis of Protein-Ligand Interactions. 9th Advanced in silico Drug Design workshop 2026. Olomouc, Czech Republic. 26th-30th January 2026.
-
Gimeno A. PDB-CAT. Invited Lecture at 9th Advanced in silico Drug Design workshop 2026. 26th January 2026.
-
Llop-Peiró A, Trujillo-De León S, Pujadas G, Garcia-Vallvé S, Gimeno A. PDB-CAT: A user-friendly tool to classify and analyze PDB protein-ligand complexes. Protein Sci. 2025 Dec;34(12):e70379. doi: 10.1002/pro.70379.
-
Fadlallah S, Serratosa F, Julià C. Graphormer Boosted: Molecular Property Prediction With Enhanced Graph Spatial and Edge Encodings. IEEE Access, 2025, vol. 13, pp. 130492-130504. doi: 10.1109/ACCESS.2025.359196
-
Segura-Alabart N and Serratosa F. Graph-Based Representations of Almost Constant Graphs for Nanotoxicity Prediction. In: Brun, L., Carletti, V., Bougleux, S., Gaüzère, B. (eds) 14th IAPR-TC15 International Workshop on Graph-Based Representations in Pattern Recognition. GbRPR 2025. Caen, France, 25-27 June 2025. Lecture Notes in Computer Science, vol 15727. Springer, Cham. doi: 10.1007/978-3-031-94139-9_6
-
Fadlallah S, Julià C, Serratosa F, Liu X, Murata T. Molecular Structure Learning with Graph Transformers: A Graph Reduction Approach for Improved Efficiency and Accuracy. 2025 IEEE 6th International Conference on Pattern Recognition and Machine Learning (PRML), Chongqing, China, 2025, pp. 411-415, doi: 10.1109/PRML66062.2025.11159676.
-
Llop-Peiró A, Garcia-Vallvé S, Ginex T, Luque J, Gimeno A, Pujadas G. Exploring SARS-CoV-2 Main Protease binding site with pharmqsar. AI in Drug Discovery and Biomedicine. Barcelona, Spain, 31 March – 2 April 2025
-
Serratosa F. AE+GAE: a mixture model for structural and semantic feature detection in graph neural network regression. Pattern Recognition Letters. 2025 195:80-86. doi:10.1016/j.patrec.2025.04.032.
-
Serratosa, F. (2025). A Generative Algorithm to Compute NanoFingerprints. In: Hernández-García, R., Barrientos, R.J., Velastin, S.A. (eds) Progress in Pattern Recognition, Image Analysis, Computer Vision, and Applications. CIARP 2024. Lecture Notes in Computer Science, vol 15369. Springer, Cham. doi: 10.1007/978-3-031-76604-6_7
-
Segura-Alabart, N., Fernández, A., Kensert, A., Cabooter, D., Serratosa, F. (2025). Evaluation Metrics in Saliency Maps Applied to Graph Regression. In: Torsello, A., Rossi, L., Cosmo, L., Minello, G. (eds) Structural, Syntactic, and Statistical Pattern Recognition. S+SSPR 2024. Lecture Notes in Computer Science, vol 15444. Springer, Cham. doi: 10.1007/978-3-031-80507-3_7
-
Serratosa F, Segura-Alabart N. MO-NanoDatabase: A metal-oxide nanostructured compound dataset composed of a huge number of metal-oxide nanocompounds, their global properties and 3D-structure. Data in Brief. 2025. 60:111476. doi:10.1016/j.dib.2025.111476.
-
Serratosa F. Graph Regression Based on Autoencoders and Graph Autoencoders. In: Antonacopoulos A, Chaudhuri S, Chellappa R, Liu CL, Bhattacharya S, Pal U (eds) Pattern Recognition. ICPR 2024. Lecture Notes in Computer Science, vol. 15310, pp 345-360. Springer, Cham 2025. doi:10.1007/978-3-031-78192-6_23
-
Garcia-Segura P, Llop-Peiró A, Novau-Ferré N, Mestres-Truyol J, Saldivar-Espinoza B, Pujadas G, Garcia-Vallvé S. SARS-CoV-2 main protease (M-pro) mutational profiling: An insight into mutation coldspots. Comput Biol Med. 2024 Nov 11;184:109344. doi: 10.1016/j.compbiomed.2024.109344.
-
Llop-Peiró A, Luque J, Ginex T, Garcia-Vallvé S, Pujadas G. Exploring SARS-CoV-2 Main Protease Binding Site with PharmQSAR. IQTC Meeting 2024. Barcelona, Spain, 30-31 October 2024
-
Llop-Peiró A, Luque J, Ginex T, Garcia-Vallvé S, Pujadas G. Exploring SARS-CoV-2 Main Protease Binding Site with PharmQSAR. 24th European Symposium on Quantitative Structure-Activity Relationship (EuroQSAR). Barcelona, Spain, 22-26 September 2024
-
Serratosa F. Graphfingerprint: graph embedding of graphs with almost constant sub-structures. Pattern Analysis and Applications 2024. 27:143. doi: 10.1007/s10044-024-01366-w
-
Serratosa, F. (2024). Graph Embedding of Almost Constant Large Graphs. In: Vasconcelos, V., Domingues, I., Paredes, S. (eds) Progress in Pattern Recognition, Image Analysis, Computer Vision, and Applications. CIARP 2023. Lecture Notes in Computer Science, vol 14469. Springer, Cham. doi: 10.1007/978-3-031-49018-7_2
-
Llop-Peiró A, Macip G, Garcia-Vallvé S, Pujadas G. Are protein-ligand docking programs good enough to predict experimental poses of noncovalent ligands bound to the SARS-CoV-2 main protease? Drug Discov Today. 2024 Oct;29(10):104137. doi: 10.1016/j.drudis.2024.104137
-
Llop-Peiró A, Pujadas G, Gimeno A, Garcia-Vallvé S. PDB-CAT: A User-Friendly Tool to Classify and Analyze PDB Protein-Ligand Complexes. ChemRxiv. 2024; doi:10.26434/chemrxiv-2024-54073
-
Llop-Peiró A, Pujadas G, Garcia-Vallvé S. PDB-CAT: Classification and Analysis Tool for PDB files. Chemoinformatics Summer School. Strasbourg, France, 24-28 June 2024
-
Llop-Peiró A, Macip G, Garcia-Vallvé S, Pujadas G. Enhancing Virtual Screening Strategies: Evaluating AutoDock Vina and Glide Methods for Predicting Conformations of Non-covalent Inhibitors Against SARS-CoV-2 Main Protease. European Workshop in Drug Design (EWDD24). Siena, Italy, 19-23 May 2024
-
Llop-Peiró A, Pujadas G, Garcia-Vallvé S. Challenges in distinguishing functional proteins from polyproteins in databases: implications for drug discovery. Brief Bioinform. 2024 Jan 22;25(2):bbae012. doi: 10.1093/bib/bbae012.
- Llop-Peiró A, Pujadas G, Garcia-Vallvé S. PDB-CAT: Classification and Analysis Tool for PDBx/mmCIF. 7th Advanced in Silico Drug Design workshop 2024. Olomouc, Czech Republic. 29th January - 1st February 2024

